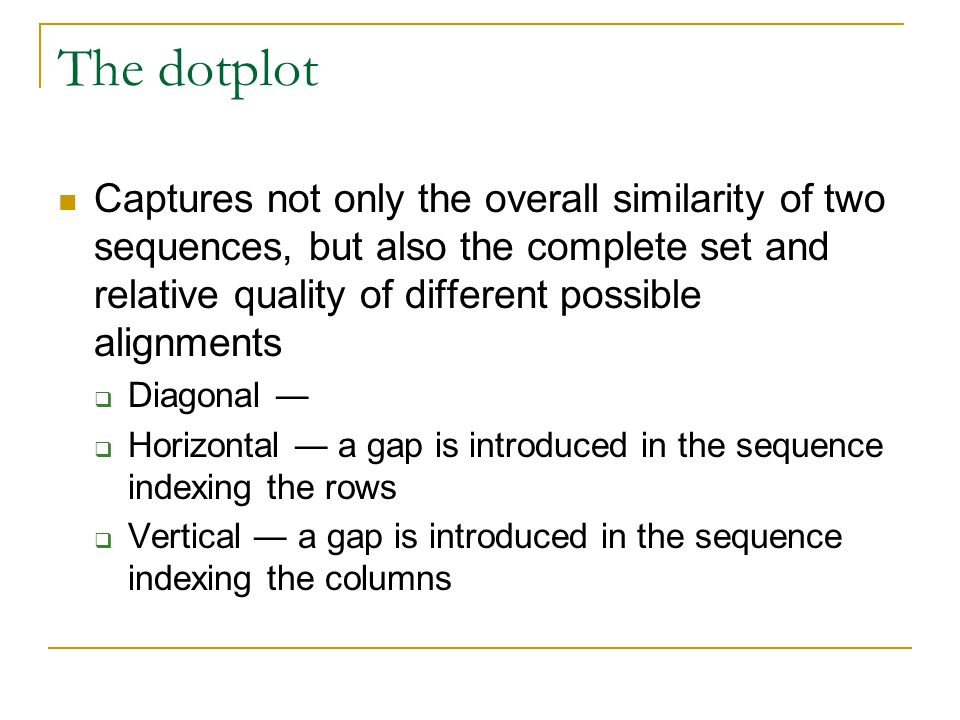
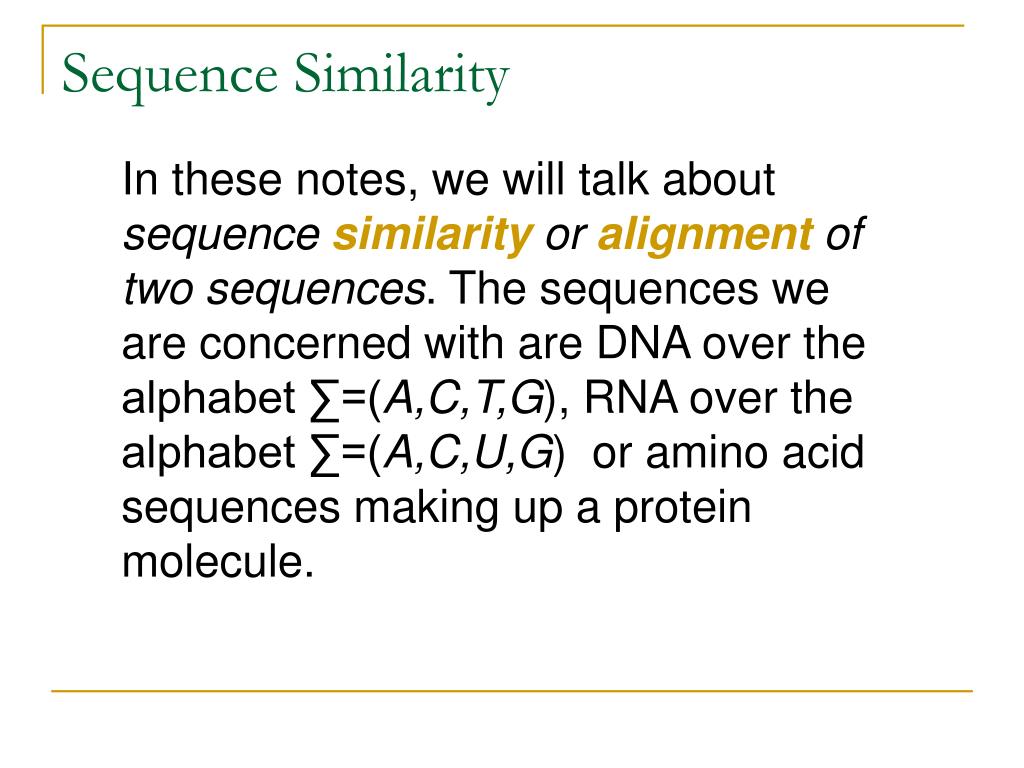
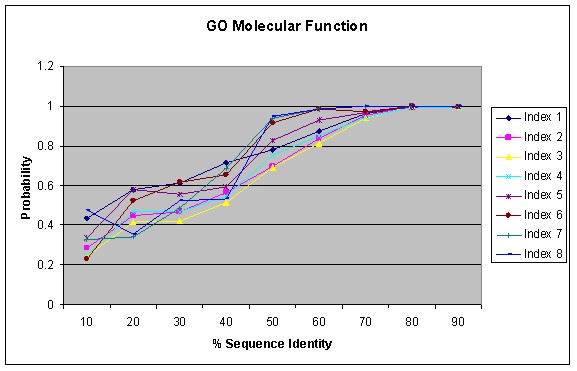
BLAST: Compare & identify sequences - NCBI Bioinformatics Resources: An Introduction - Library Guides at UC Berkeley
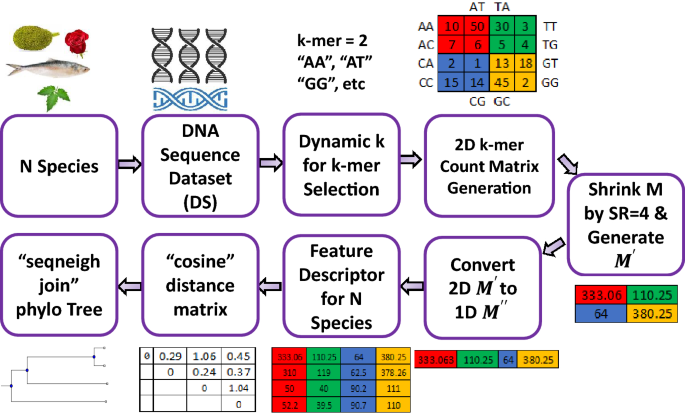
A fast and efficient algorithm for DNA sequence similarity identification | Complex & Intelligent Systems

SOLVED: What is a multiple sequence alignment good for? a. To identify conserved and varying residues (i.e. base or amino acid) in similar sequences. b. To infer functional roles to proteins and
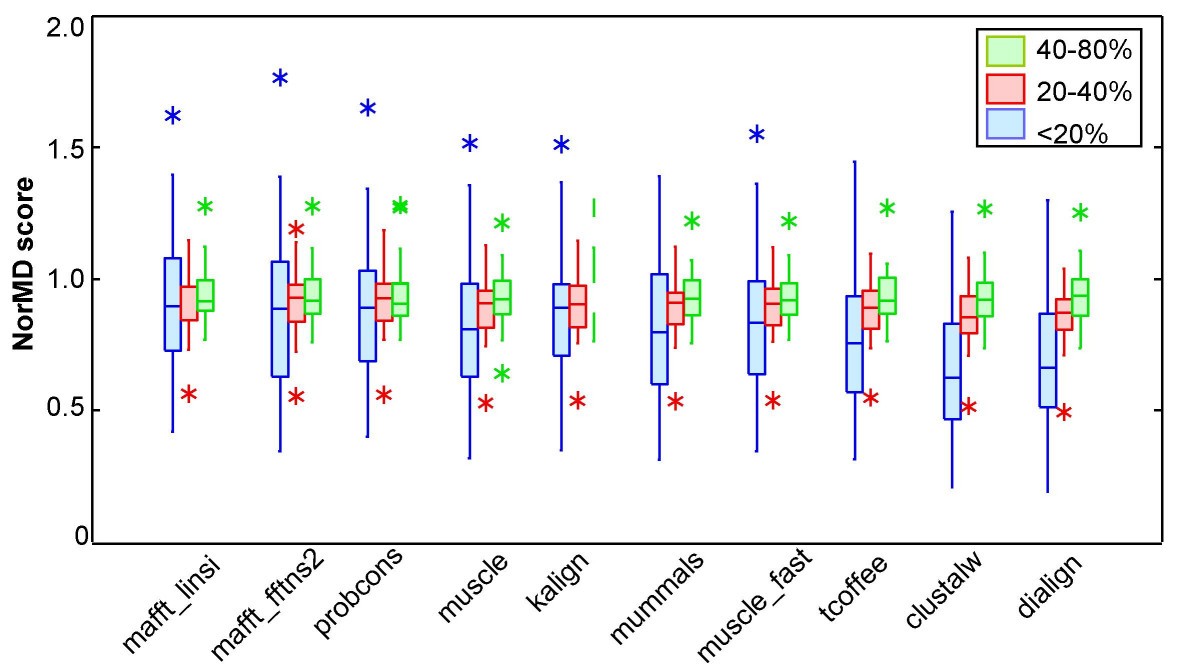
A new protein linear motif benchmark for multiple sequence alignment software | BMC Bioinformatics | Full Text
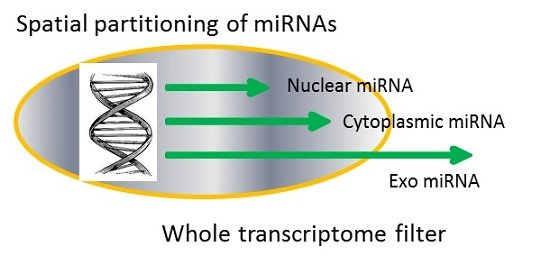
IJMS | Free Full-Text | Spatial Partitioning of miRNAs Is Related to Sequence Similarity in Overall Transcriptome

A) 16S rRNA gene sequences similarity and overall genome relatedness... | Download Scientific Diagram
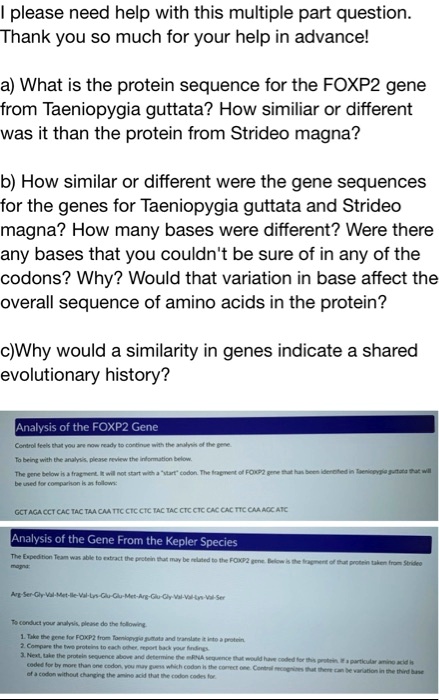

SOLVED: please need help with this multiple part question. Thank you so much for your help in advancel a) What is the protein sequence for the FOXPZ gene from Taeniopygia guttata? How

Difference Between Similarity and Identity in Sequence Alignment | Compare the Difference Between Similar Terms

Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences | PNAS

BLAST: Compare & identify sequences - NCBI Bioinformatics Resources: An Introduction - Library Guides at UC Berkeley

1.2: Identifying Conserved Elements in the Toxin Sensor and Designing Mutants to Test Whether They are Important for Function - Biology LibreTexts
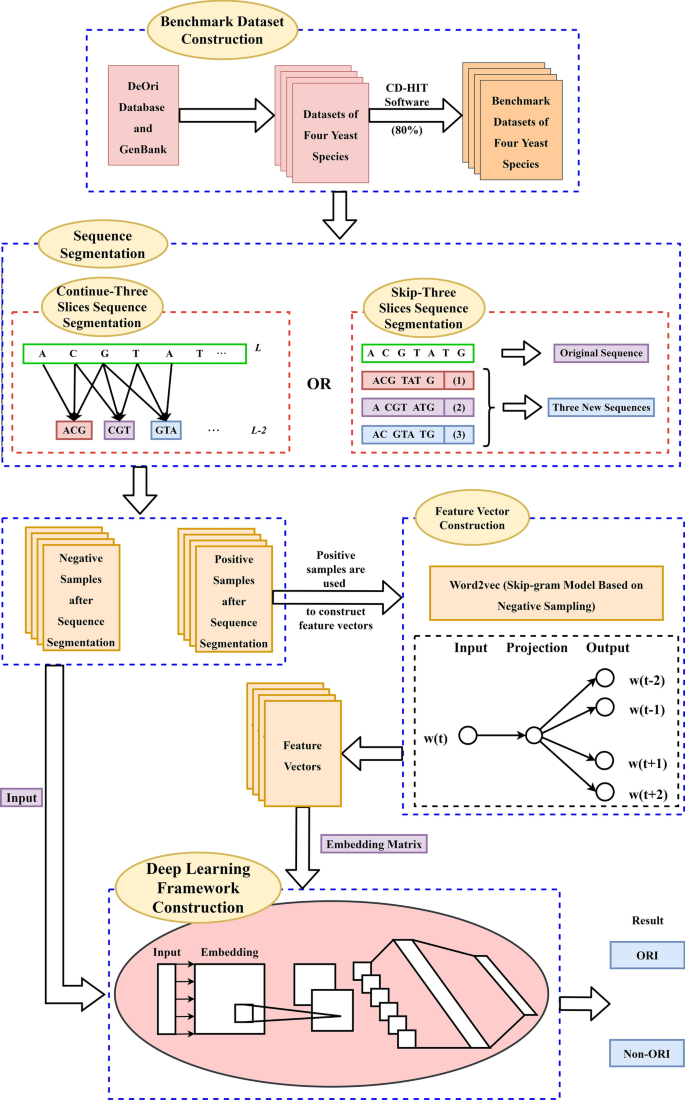
A deep learning framework combined with word embedding to identify DNA replication origins | Scientific Reports
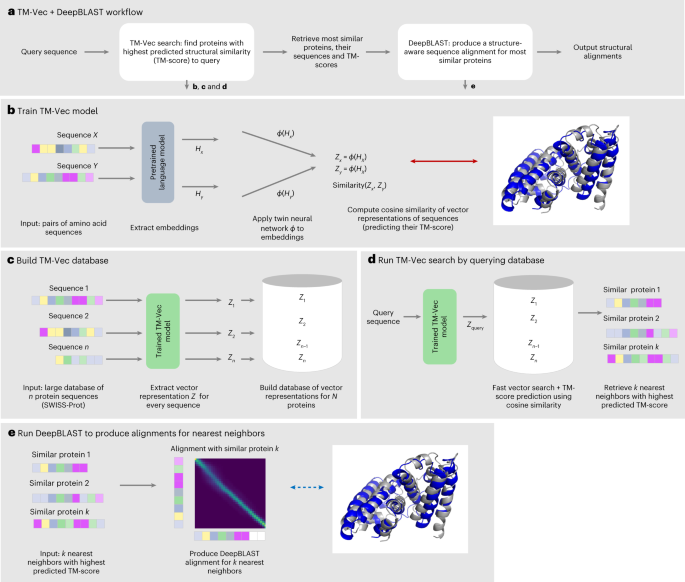
Protein remote homology detection and structural alignment using deep learning | Nature Biotechnology

SRSCL: A strong-relatedness-sequence-based fine-grained collective entity linking method for heterogeneous information networks - ScienceDirect